Signed in as:
filler@godaddy.com
Signed in as:
filler@godaddy.com
These cells are coming from mouse pancreas epithelium, transformed to express a mutant form of RAS. One of their characteristic features is prominent nucleoli that sometimes arrange to make little smiley faces.
Depicted is a group of extracellular vesicles (EVs) derived from iPSC-microglia. The largest contains fibrils believed to be tau, a protein that misfolds in several neurodegenerative diseases. Microglia ingest tau and attempt to degrade it but may package it into EVs for release. .
Xist RNA dynamics within mESC nuclei

Enchanted ciliary forest by Elisa Vitiello, Gergely lab, Department of Biochemistry.
In this image, we can observe tiny cilia of retinal pigmental epithelial cells (RPE1cells). In green, acetylated tubulin, main component of the cilia axoneme and centriole, foundation of the antenna. These tiny cilia, like antennas, perceive signals from the external environment to coordinate cellular responses.

Kind Wishes by Zeinab Rekad, Mardakheh Lab, Department of Biochemistry.
These cells are coming from mouse pancreas epithelium, transformed to express a mutant form of RAS. One of their characteristic features is prominent nucleoli that sometimes arrange to make little smiley faces.

A partridge in a (Mismatch) Repair tree by Amy Moores, Uphoff Lab, Department of Biochemistry.
The image shows real single-molecule trajectories of fluorescently labelled E.coli mismatch repair enzymes that have been arranged into the shape of a Christmas tree. We have used a bright stable fluorescent dye called JFX650 for labelling, allowing us to follow the same individual enzyme around the cell

The Christmas Tree in the Cerebellum by Yin Chun Cheng, Becker lab, NDCN.
Purkinje cells are basically biological Christmas trees. Located in the cerebellum, their huge dendritic branches are critical for motor coordination, but here they look festive. This Airyscan image captures every detail, turning the tiny synaptic spines into glowing ornaments on the boughs. I’m dedicating this to Jan Evange

A Festive Burst of Cellular Architecture by Elisa Vitiello, Gergely lab, Department of Biochemistry.
We uncovered a new role for EDC4, previously thought to function only in RNA regulation. Beyond maintaining RNA stability, EDC4 also plays a key role in building primary cilia, the “antennae” cells use to sense their environment. linked to Parkinson’s disease, our findings suggest broader roles in

Nucleic Acid Snowman by Isabella Savin, Schermelleh lab, Department of Biochemistry.
My image depicts a snowman in a flurry of snow, created from 3D-SIM and widefield images of DNA and RNA staining in C127 cells. The snowman wears a scarf and hat made of nucleoli, where ribosomal RNA is produced. Inspired by a childhood memory of building a snowman in my grandparents’ garden, I dedicate this image

NeuroWreath by Nick Gatford, Baldwin lab, Department of Chemistry.
A festive wreath of individual human neurons on a wintry background of synaptic vesicle protein staining, with the red ribbons of a protein structure of interest as a bow.

Xist-MAS Tree 🎄🎄🎄 by Adam Cawte, Brockdorff lab, Department of Biochemistry.
Xist-RNA in Green
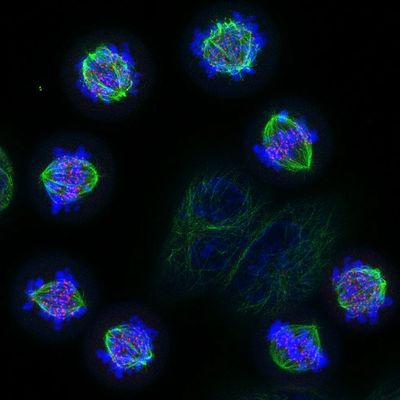

We are thrilled to celebrate Dr. Nick Gatford on his recent achievement in the NIKON Small World competition, earning a prestigious Image of Distinction award! This year’s competition, which marks the 50th anniversary of incredible images captured by the light microscope, showcased stunning visual breakthroughs from around the world, and we’re proud to see Nick’s work recognised at such a renowned level. His accomplishment highlights the power of microscopy in revealing the unseen beauty of the world around us, and it's inspiring to everyone in our Micron community.
The winning image shows a tiled network of dopaminergic neurons generated from human stem cells. Dopaminergic neurons are the main cell type that deteriorates in Parkinson’s disease. We use these cells to discover new pathways to prevent this degeneration. A colour encoded depth-based filter highlights the extent to which these neurons form an interactive network.
Human iPSCs carrying a triplication in the synuclein gene were differentiated into human dopaminergic neurons by applying various small molecules that mimic the signals associated with dopaminergic cell differentiation in mammalian neurodevelopment. Neurons were differentiated for 80 days at which point they were fixed and stained for MAP2 to identify neurons and two further proteins of interest that are key to Parkinson’s disease pathogenesis. The image displays MAP2 staining to identify cells appropriate for further examination and was acquired using a 60X objective attached to a Zeiss Airyscan LSM980 microscope in the Micron facility. The image was post-processed in ImageJ where a colour encoded depth-based filter was applied to the complete 10x10 tiled set of z-stacks, consisting of approximately 8,000 individual images projected in z to show the entire network in this combined field of view.

The new Stellaris TauSTED Xtend has now been installed at Micron as part of the ongoing Leica Centre of Excellence collaboration.
"TauSTED Xtend, enables extended multicolor imaging of live and intact specimen at nanoscale. By combining spatial and lifetime information, TauSTED Xtend gives access to an extra level of information that allows resolving small details in context at extremely low light dose" - Leica.
Multicolor fixed STED image. Image courtesy of Dennis Derstrof, Klinik für Hals-Nasen und Ohrenheilkunde, Universität Marburg & Prof. Dr. Dominik Oliver aus dem Institut für Physiologie und Pathophysiologie, Abteilung für Neurophysiologie, Universität Marburg.

A VT-iSIM system has recently been installed at Micron & will soon be available for access. Funded by the ERC, John Fell & EPA Cephalosporin Fund.
The system is equipped for live cell super-resolution imaging with twin Hamamatsu Orca Quest qCMOS cameras.
Image left shows F-actin activity in human neuronal growth cones.
iSIM images by Nick Gatford, Nuffield Department of Clinical Neurosciences.
The iSIM was used to show that localised clustering of autism spectrum disorder (ASD)-associated proteins NLGN3/4X at the leading edge of growth cones promoted neuritogenesis in immature human neurons by inducing growth cone enlargement and influencing actin filament organization. Ultimately suggesting these proteins may have a role in earlier ASD pathogenesis prior to their established role at neuronal synapses. The VT-iSIM enabled visualisation of these key proteins of interest at the nanoscopic level in a human stem cell derived neuronal system for the first time, and provided unprecedented detail at speed to generate a substantial dataset of super-resolution images suitable for quantification and publication https://pubmed.ncbi.nlm.nih.gov/34542148/.

Niloufer Irani and Deirdre Kavanagh have both been nominated as one of many inspirational women in the Department of Biochemistry as part of the department's International Women's Week celebrations.
"I would like to nominate Deirdre because she is always willing to help, she is an incredible teacher, and she is just generally a lovely warm person! Her expert knowledge of microscopy techniques and her amazing problem-solving skills means that she always gives us the best advice for helping us to achieve our results... Our department is so lucky to have her!"
The official inauguration of the Leica Centre of Excellence at the Micron Bioimaging Facility, Department of Biochemistry will be held on September 22nd. Through this strategic partnership researchers will have access to the latest advances in microscopy from Leica hosted at Micron including confocal, super-resolution, FLIM and STED capabilities.
Location
Department of Biochemistry
Seminar Room 20-138 Ground Floor
Dorothy Crowfoot Hodgkin Building
Schedule of the day
10 am Welcome addresses from Prof Francis Barr, Associate Prof Lothar Schermelleh, Department of Biochemistry and Darin Stell, Senior VP, Global Commercial Operations, Leica
10:30 am Mica: Step into the Microhub Era - Dr. Annalisa Rizza, Leica Microsystems
10:55 am Correlative Light and Electron Microscopy at COSMIC and Micron - Dr Lindsay Baker, Kavli Institute for Nano discovery
11:05 am The Power to See More with the Stellaris Confocal Platform - Paul McCormick, Leica Microsystems
11:30 am Past, present and future of Lifetime imaging - Dr Sergi Padilla Parra, Department of Infectious Diseases, Kings College London
12:30 pm Official ribbon cutting and lunch
13:30 pm Microscopy showcase of the Mica and Stellaris 5 at Micron Bioimaging Facility
To register for the launch and to book a demo of the Mica or Stellaris 5 please use the registration button below.

The Department of Biochemistry's Centenary Scientific Imaging Competition was held to celebrate a hundred years of research excellence.
The competition, sponsored by ONI, attracted entries from students, researchers, and staff members across the MSD, all showcasing their scientific imaging skills.
How long have you been with Biochemistry and where did you work before that ?
I joined the Department of Biochemistry in May 2021 as manager of the Micron Bioimaging Facility. Prior to this I worked as the lead imaging specialist at the Centre of Membrane Proteins and Receptors (COMPARE) at the University of Birmingham. I joined COMPARE after doing two post-docs in Edinburgh where I applied super-resolution microscopy to study cell communication. During this time, I started to look for opportunities outside of my own research career as I wanted to help the broader scientific community. My PhD is in Engineering and Physical Sciences and not biology!
Job title: Facility Manager, Micron Bioimaging Facility.
What are the main elements of your role?
My job is multifaceted, involving both managerial and scientific expertise. A typical day can involve training and teaching researchers in advanced microscopy, collaborating and advising on research projects, designing experiments, maintaining the facility labs and equipment, writing grants, research papers, risk assessments, managing the facility finances and developing facility strategies. The role also involves organising events and courses, testing new technologies, maintaining our website (www.micronoxford.com) and social media profile (@MicronOxford). I really enjoy the diversity and working directly with people across projects that have impact in many fields.
What’s the most common question you get asked as part of your role?
During project discussion meetings, I frequently get asked “Which microscope technique should I use?” This an important question and the answer really depends on what biological question you are trying to answer. In general, if you have never even looked at your sample before using a fluorescent microscope, then you are going to start with a confocal or widefield microscope. Anything else is going to over complicate your experiments and potentially waste a lot of time. If you are interested in live cell dynamics then widefield microscopy is the go-to as it is much quicker and will induce less phototoxicity in your sample. This can then be followed up with deconvolution to reduce out of focus light, a characteristic of widefield imaging. A particular strength of ours in Micron is in super-resolution microscopy. These techniques are extremely powerful and will allow you to view events or structures under the diffraction limit. There are many different flavours of super-resolution microscopy (SIM, SMLM, STED etc) and which one you use again depends on your biological question. Please do get in contact (micron@bioch.ox.ac.uk) and we would be very a happy to discuss all the options available in Micron.
Favourite film: I love science-fiction, from Inner Space (1987) to Interstellar (2014)
Favourite book: I have two: Martel’s Life of Pi, and du Maurier’s Rebecca
Favourite album: Fleetwood Mac – Rumours
Favourite place to go on holiday: Home to friends and family, in Ireland and Italy
Favourite place to eat in Oxford: I am still new to Oxford, so I am looking forward to finding my favourite place here. In fact, I would be happy to receive suggestions for the best pizzeria around!
What do you like to do when you’re not at work: I like to do gardening mainly to grow my own fruit and veg. Plus: cycling, visiting museums and botanical gardens, going wine tasting, and planning my next holiday (which will likely include all of the above).


The 7th summer school of the Edinburgh Super-Resolution Imaging Consortium (ESRIC) took place from 8th – 16th June 2021. Combining lectures from experts in the field with online workshops, the course featured a Tips and Tricks SMLM workshop from Micron facility manager Deirdre Kavanagh.
Out now
The Theo Murphy meeting issue‘Super-resolution structured illumination microscopy’ organised and edited by Kirti Prakash, Benedict Diederich, Stefanie Reichelt, Rainer Heintzmann and Lothar Schermelleh. Available here.
Cover image credit: Lothar Schermelleh. Wide-field vs. super-resolution 3D-SIM image of EdU labelled DNA replication units in a DAPI stained mouse cell nucleus, with corresponding spatial frequency distribution below.

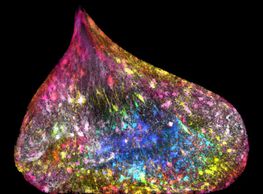
Congratulations to Ezequiel Miron from the Schermelleh Group, Department of Biochemistry at the University of Oxford whose image is one of the selected winners of the Wellcome Image Awards 2017.
His picture, captured using the OMX V3 super-resolution microscope in Micron, shows the nucleus of one of two new daughter cells. The DNA in this cell has somehow become caught, and is being pulled between the two dividing cells causing the DNA to be stretched out inside the nucleus; DNA fibres can be seen running through it. As the new cells have moved apart the DNA is being pulled through a perforation in the nuclear membrane. The tension distributed by the rope-like DNA has deformed the usually circular envelope of the nucleus.

On the 25th and 26th of June, Micron was involved in setting up one of thirty stalls at the Oxford Science Fair, part of the Oxford Science Festival 2016. We joined scientists from across Oxford to exhibit work from neurology, quantum computers and even the future of our diet (FYI… its insects!); we, of course, brought the microscopes! Our stall was a joint effort with members of the Srinivas and Mommersteeg groups in the Department of Physiology, Anatomy & Genetics, focusing on developmental biology. The stall was packed with chick and zebrafish embryos and dissected mouse hearts at different stages of their development, as well as live zebrafish embryos under a microscope connected to a screen via a Raspberry Pi camera from Micron. Children and adults alike gasped wide-eyed at the sight of the beating heart on screen, visible through the transparent body of the zebrafish. As further visual aids we projected movies and microscope time-lapse experiments of the development of zebrafish, chick, mouse and human embryos.
Ezequiel Miron from the Schermelleh group at Micron took the data used in making the 3D printed models to generate explorable giant scale replicas within the world of the game Minecraft. These virtual worlds were projected on the big screen and generated a lot of interest among the young boys visiting our stall, and the parents who wanted to know where to download them for their kids to explore at home.
That achievement along with many other smiling faces made this event a wonderful experience in public engagement.
The Minecraft worlds of the developing heart and the mammalian nucleus can be found at:http://www.schermellehlab.org/minecraft/
Many more pictures of this event can be found at: http://www.schermellehlab.org/oxsci/

Micron has been awarded a renewal of its Strategic Award from the Wellcome Trust to continue its state-of-the-art collaborative activities until at least 2020.
Micron's Director Ilan Davis, together with deputy-Director Jordan Raff at the Dunn School, aim to expand the co-ordinating role played by Micron in this new phase, bringing in additional departments and technologies. This includes Christian Eggeling from the WIMM, Martin Booth from Engineering, Yvonne Jones and Kay Grunewald from the WTCHG, and Achillefs Kapanidis from Physics.
The £4.6million Strategic Award will support the purchase or construction of bespoke microscopes, the development of new tools (analytical and chemical), and increase access and support for users.
The new investment, together with substantial support from the University and existing support from the MRC and other funders, will allow Micron to continue to develop as one of the UK's leading bio-imaging facilities.
'Micron is an exemplar of how complex technologies can be offered to researchers,' says Professor Davis. 'There's been a shift in biology to big technologies that cannot be supported by single labs, and the field is evolving so fast that individuals cannot keep up with this.' 'The infrastructure required to make the technologies available widely is substantial. It's not a question of simply pressing a button - for example, the data analysis aspect is complex and needs a deep understanding.'
The award will enable the development of new imaging technologies as well as the continued development of existing microscopy systems. Off-the-shelf microscopes purchased from Zeiss, which provide moderate super-resolution and are very sensitive, will expand Micron's capabilities. Two new systems, the Lattice Lightsheet system and the 4Pi-SMS microscope, will be built.
Janelia Campus Nobel Laureate Eric Betzig recently developed the revolutionary Lattice Lightsheet system (Science (2014) Vol. 346). It is ideal for rapid and non-disruptive long-term live cell imaging as it has particularly good light efficiency and therefore causes less damage. Christian Eggeling will lead this work at the WIMM where the microscope will be used to look at the immune system responses live. It will be one of the first instruments of its type in the UK.
The single-molecule high-resolution system 4Pi-SMS is tailored to deep imaging of samples. It was developed by Jim Rothman at Yale and a prototype is currently being built at the Gurdon Institute in Cambridge. In Oxford, Martin Booth in the Department of Engineering has developed the necessary adaptive optics, which allow corrections of aberrations, to image deep. He will spearhead the building of a replica microscope making Micron's instrument the third in the world after Yale and the Gurdon Institute.
The continued development of existing microscopy systems will focus on completion of a DeepSIM microscope, which has already progressed well from a long-standing collaboration with Professor John Sedat at UCSF. The system lends itself to specimen manipulation, such as microinjection, at the same time as imaging. Like 4Pi-SMS, it uses adaptive optics and there are no commercial systems currently available. Micron researchers have developed user-friendly software that will make the system easy to use for biologists.
Micron will continue its collaboration with Diamond to improve a version of structured illumination microscopy (a mode of super-resolution microscopy) to be used by researchers alongside X-ray microscopy and EM. The lab of Achillefs Kapanidis, Professor of Biological Physics at Oxford, has developed the NanoImager, a compact desktop microscopy for single molecule analyses, in collaboration with David Sherratt in Biochemistry.
Professor Davis comments that the diversity of expertise across Oxford and willingness of individuals to work together has been crucial to Micron's success. 'The landscape of researchers in Oxford is great and there is a high degree of co-operation between them. We are also fortunate to have tremendous institutional support - both financially and in support of the concept.'
The equipment at Micron is available to everyone across Oxford and where appropriate, outside Oxford.
Article by Jane Itzhaki